Evaluation of Sesame Genotypes for Field Resistance Against Stem And Root Rot Caused by Macrophomina Phaseolina (Tassi) Goid.
0 Views
B.VISHNU*, K. VISWANATH, P. NAGAMANI AND D. BHARATHI
Department of Plant Pathology, ANGRAU-S.V. Agricultural College,Tirupati-517 502.
ABSTRACT
Stem and root rot caused by Macrophomina phaseolina is a devastating disease in sesame. Use of host plant resistance is the cheapest and effective disease management strategy. In the present study, 58 sesame genotypes were evaluated for their resistance against stem and root rot disease in 2025 in a sick field under artificial conditions. The results showed that there was significant difference in Percent Disease incidence (PDI %) of the genotypes tested. The mean disease incidence of genotypes ranged from 5.7 to 59.8 %. Eight genotypes VSP-16 (5.7 %), TPTS-40 (6.6 %), YLM-161 (8.6 %), Nirmala (8.9 %) and VS-16- 009 (9.5 %), Madhavi (9.6 %), GT-10 (9.6) and VS-20-008 (10 %) were found to be resistant (R). These genotypes might be helpful in improving the breeding program of sesame crop. These sesame genotypes can be used for improvement of genetic resistance in already developed lines/varieties or development of new resistant varieties and cultivars through hybridization program.
KEYWORDS: Genotypes, percent disease incidence, resistance.
INTRODUCTION
The queen of oil seed, sesame is known for its high nutritional and therapeutic value (Biswas et al., 2018). It have been consumed as traditional healthy food for its anti-inflammatory and antioxidative properties (Yadav et al., 2022). The crops susceptibility to disease is the main barrier to the sesame production causing 7 million tons of yield loss for every year (Ara et al., 2017) and it creates significant impact on growers. Stem and root rot caused by M. phaseolina is a devastating disease in sesame and causes 5-100 per cent yield loss (Vyas, 1981). Murugesan (1978) estimated that increase in 1 per cent of disease infection may lead to yield loss of 1.8 kg ha-1.
As stem and root rot is a seed, stubble and seed borne disease, it is difficult to control by conventional management practices. Using chemicals not only increase input cost but also harmful to environment. Various management studies have been carried out by several researchers viz., using bio-agent (Abdul sattar et al., 2006), plant extracts (Ahmed et al., 2010) and cultivating resistant varieties (Thiyagu et al., 2007). Among these, host plant resistance is one of the important effective strategy for the management of disease (Salari et al., 2012). Different genotypes of sesame will differ in their genetic potential of disease resistance. The screening of genotypes for resistance against stem and root rot will help to identify genotypes for resistance.
Hence, the present research was planned out to evaluate the resistance of sesame genotypes against stem and root rot disease.
MATERIAL AND METHODS
Fifty eight sesame genotypes obtained from AICRP on sesame, Tirupati were evaluated for their resistance against stem and root rot disease under sick plot conditions during the year 2025. Each genotype was sown in a row of 3 m length with inter row spacing of 30 cm. A row of susceptible check was sown after every four genotypes. Two replications of each genotype was maintained. Two susceptible checks, YLM-66 and YLM-146 were used as infector rows. Disease incidence were recorded at 70 days and Percent Disease Incidence (PDI) was calculated.
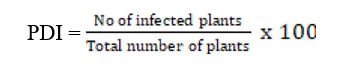
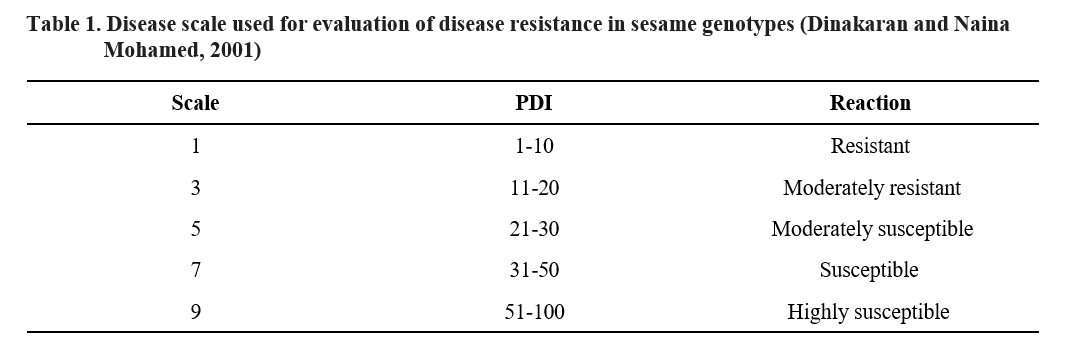
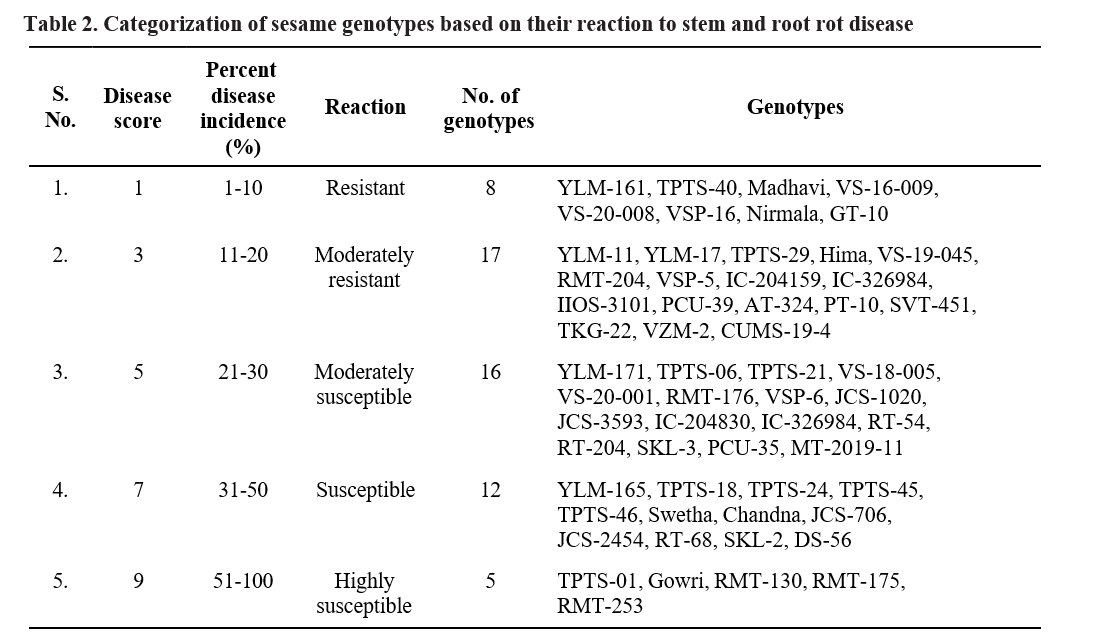
Based on PDI, the genotypes were categorized as resistant, moderately resistant, moderately susceptible, susceptible and highly susceptible (Table 1).
RESULTS AND DISCUSSION
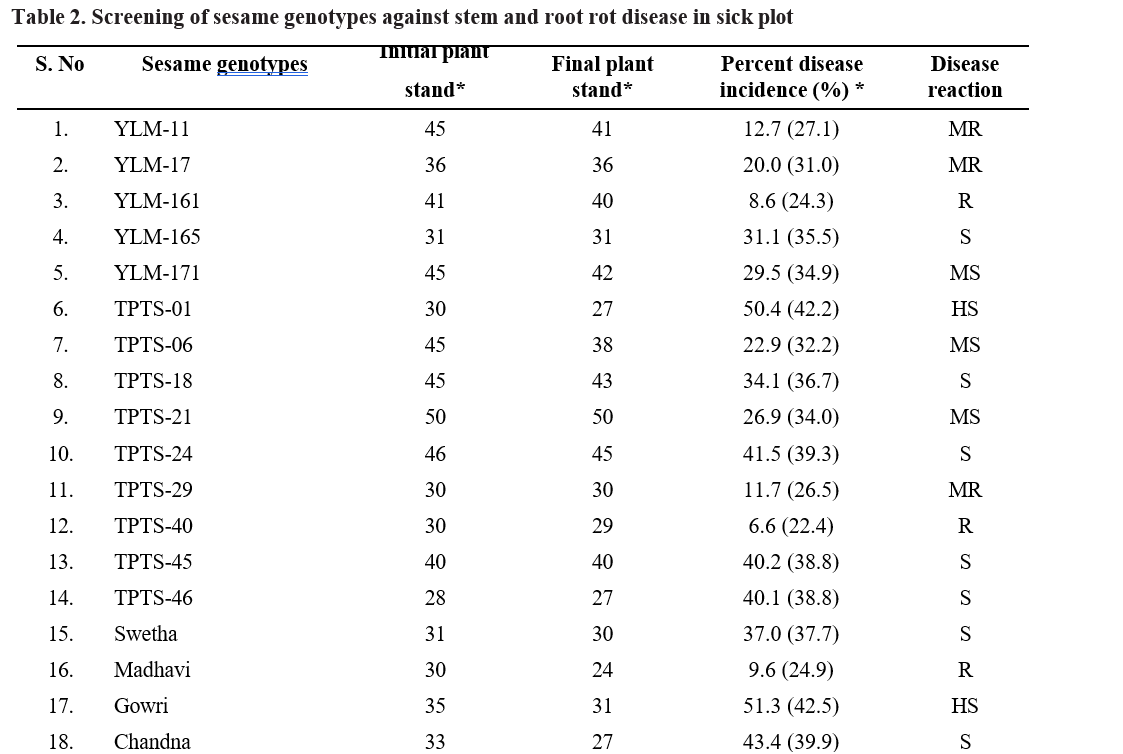
Results from the field experiment on screening of sesame genotypes against stem and root rot revealed that mean disease incidence of genotypes ranged from
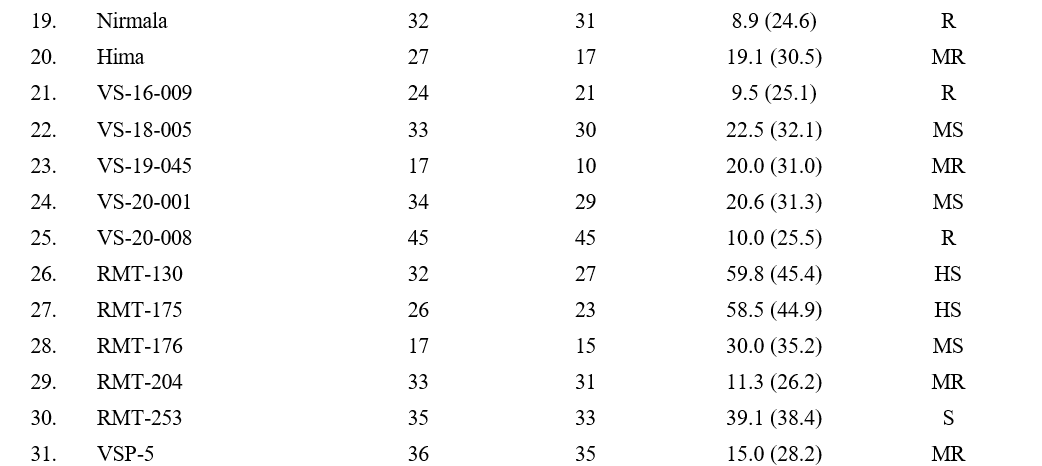
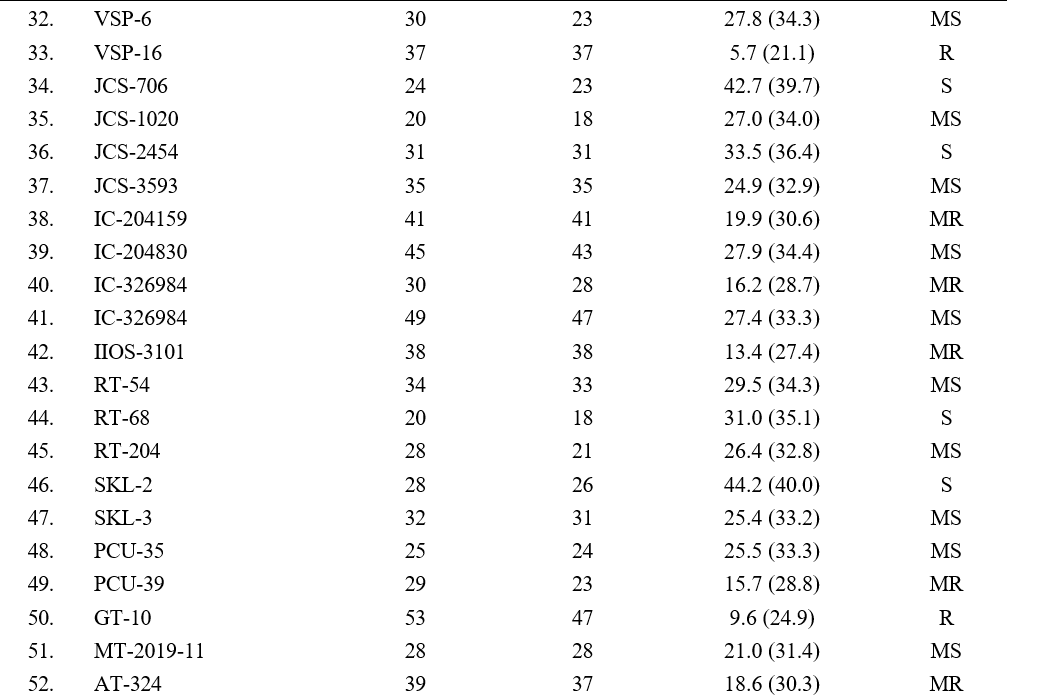
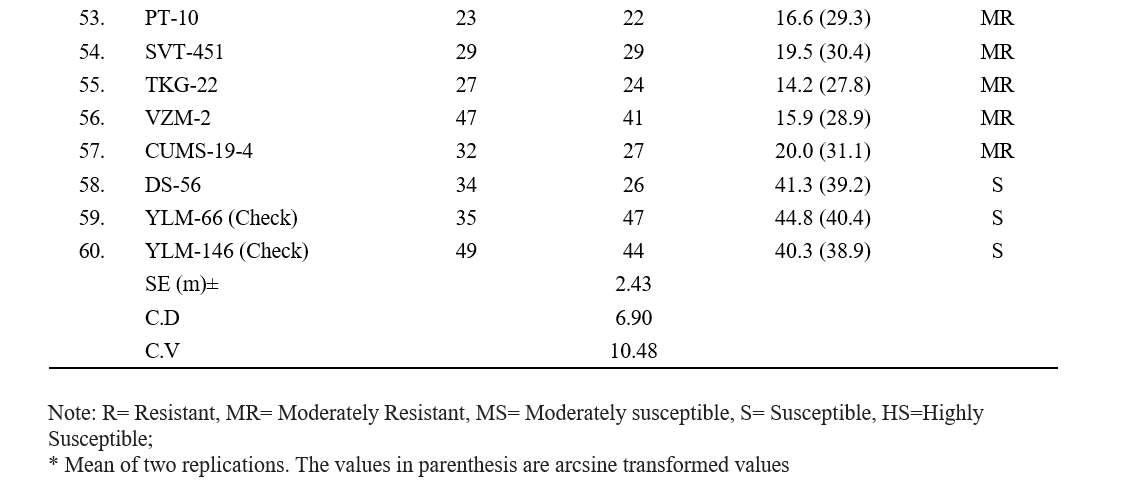
5.7 to 59.8 per cent (Table 2). The least disease incidence was observed in genotype VSP-16 with 5.7 per cent, followed by genotypes TPTS-40 (6.6 %), YLM-161 (8.6 %), Nirmala (8.9 %) and VS-16-009 (9.5 %). The highest disease incidence was observed in genotype RMT-130 with 59.8 % followed by RMT-175 (58.5), Gowri (51.3 %), TPTS-01 (50.4) and SKL-2 (44.2 %). Remaining genotypes (48) showed disease incidence ranging from 9.6 to 43.4 per cent.
Based on the per cent disease incidence, out of 58 genotypes screened, 8 genotypes (YLM-161, TPTS-40, Madhavi, VS-16-009, VS-20-008, VSP-16, Nirmala, GT-10) were found to be resistant (R) with >10 per cent disease incidence, 17 genotypes (YLM-11, YLM- 17, TPTS-29, Hima, VS-19-045, RMT-204, VSP-5, IC-204159, IC-326984, IIOS-3101, PCU-39, AT-324, PT-10, SVT-451, TKG-22, VZM-2, CUMS-19-4) were found moderately resistant with 11-20 per cent disease incidence, 16 genotypes (YLM-171, TPTS-06, TPTS- 21, VS-18-005, VS-20-001, RMT-176, VSP-6, JCS- 1020, JCS-3593, IC-204830, IC-326984, RT-54, RT-204, SKL-3, PCU-35, MT-2019-11) were found moderately susceptible with 21-30 percent disease incidence, 12 genotypes (YLM-165, TPTS-18, TPTS-24, TPTS-45, TPTS-46, Swetha, Chandna, JCS-706, JCS-2454, RT- 68, SKL-2, DS-56) showed susceptible reaction with 31-50 percent disease incidence and 5 genotypes (TPTS- 01, Gowri, RMT-130, RMT-175, RMT-253) were found to be highly susceptible with disease incidence of more than 51 per cent (Table 3).
Similarly, the disease reaction of sesame genotypes to stem and root rot disease was studied by Thiyagu et al. (2007). In sick plot method, fifteen parents and their F1’s were screened against the disease. Three genotypes (ORM 7, ORM 14 and ORM 17) were found to be resistant with PDI of 7.69, 8.33 and 6.67 respectively in sick plot condition but the F1’s screened against the disease showed that all crosses were susceptible to highly susceptible to root rot disease. Ramesh Nath Gupta et al. (2020) evaluated three hundred fifty germplasms lines for their relative resistance under natural sick plot conditions and majority of them showed moderately susceptible to susceptible reaction against stem and root rot. None of the germplasms showed resistant reaction. Similar studies by Deepthi et al. (2014) revealed that none of the cultivars showed complete resistance against phaseolina. Only one entry PKDS-91 was found as moderately resistant to charcoal rot. Ezhilarasi and Meena (2019) found most of the sesame lines were moderately susceptible to stem and root rot in their study.
Even though many lines have been identified as moderately resistant against the stem and root rot disease, complete resistant genotype are rarely available against this wide host range pathogen. In the study, eight genotypes showed resistant disease reaction against the stem and root rot. But further evaluation of these genotypes in different seasons under variable conditions will reveal the resistant potential. Hence resistant genotypes are recommended for further evaluation and potential cultivation. These lines might be helpful in improving the breeding program of sesame crop. These sesame lines can be used for improvement of genetic resistance in already developed lines/varieties or development of new resistant varieties and cultivars through hybridization program.
The identification of resistant genotypes in the study marks a significant step toward sustainable management of stem and root rot in sesame. Continued
efforts in screening, validation and breeding will contribute to securing sesame production and improving farmer livelihoods. The use of resistant varieties not only provides an eco-friendly and cost-effective management strategy but also reduces the dependency on chemical control measures.
LITERATURE CITED
Abdul Sattar, M.H., Salem, A.M and Bayounes, A.A. 2006. Physical and biological control of charcoal root rot in sesame caused by Macrophomina phaseolina (Tassi) Goid. Arab Journal of Plant Protection. 24: 37-40.
Ahmed, H.A.M., Abdel Razik, A.A., Hassan, M.H.A. and Khaled, S.A. 2010. Management of charcoal rot of sesame by seed soaking in medicinal plant extracts and hot water. Plant Pathology Journal. 26: 372 379.
Ara, A., Akram, A., Ajmal, M., Akhund, S., Nayyar, B.G., Seerat, W and Chaudhry, S.M. 2017. Histopathological studies of sesame (Sesamum indicum) seedlings infected with Fusarium oxysporum. Plant Pathology & Quarantine. 7: 82 90.
Biswas, S., Natta, S., Prasad Ray, D., Mondal, P and Saha, U. 2018. Til (Sesamum indicum L.) – An Underexploited but Promising Oilseed with Multifarious Applications: A Review. International Journal of Bioresource Science. 5: 127-139.
Deepthi, P., Shukla, C.S., Verma, K.P and Reddy, E.S.S. 2014. Identification of charcoal rot resistant lines of Sesamum indicum and chemical management of Macrophomina phaseolina. Medicinal Plants. 6(1): 36-42.
Dinakaran, D and Naina Mohammed, S.E. 2001. Sesame and Saffiower Newsletter. 16: 68-71.
Ezhilarasi and Meena. 2019. Screening for resistance to Macrophomina root rot in advanced breeding lines of sesame. Journal of Plant Development Sciences. 11(5): 303-305.
Murugesan, M., Shanmugam, N., Menon, P.P.V., Arokiaraj, A., Dhamo, K.P. and Kochubabu, M. 1978. Statistical arrangement of yield loss of sesame due to insect pests and diseases. Madras Agriculture Journal. 65: 290-295.
Ramesh Nath Gupta, Saharan, H. S., Rathi, A.S and Ram Avtar. 2020. Identification of Resistance Sources against Charcoal Rot of Sesame Incited by Macrophomina phaseolina (Tassi) Goid. International Journal of Current Microbiology and Appied Science. 9(2): 1795-1801. doi: https://doi. org/10.20546/ijcmas.2020.902.205
Salari, M., Panjehkeh, N., Nasirpoor, Z and Abkhoo, 2012. Screening of Cucumis melo L. Cultivars from Iran for Resistance against Soil-Borne Fungal Pathogens. Journal of Plant Pathology and Microbiology. 3(6). doi:10.4172/2157 7471.1000138.
Thiyagu, K., Kandasamy, G., Manivannan, N., Muralidharan, V. and Manoranjitham. S.K. 2007. Identification of resistant genotypes to root rot disease (Macrophomina phaseolina) of sesame (Sesamum indicum L.). Agricultural Science Digest. 27: 34 – 37.
Vyas, S.C. 1981. Diseases in sesame in India and their control. Pesticides. 15: 9-10.
Yadav, R., Kalia, S., Rangan, P., Pradheep, K., Rao, G.P., Kaur, V., Pandey, R., Rai, V., Vasimalla, C.C., Langyan, S., Sharma, S., Thangavel, B., Rana, V.S., Vishwakarma, H., Shah, A., Saxena, A., Kumar, A., Singh, K and Siddique, K.H.M. 2022. Current Research Trends and Prospects for Yield and Quality Improvement in Sesame, an Important Oilseed Crop. Frontiers in Plant Science. 13: 863521. doi: 10.3389/fpls.2022.863521
- A Study on Profile of Farmers on Drone Technology Application in Agriculture of Rayalaseema Region of Andhra Pradesh
- Studies on Combining Ability for Grain Yield and Its Attributing Traits In Maize (Zea Mays L.)
- Aphidophagous Coccinellid Fauna Associated With Pulses and Oilseed Crop Ecosystems in Tamil Nadu
- Evaluation of Sesame Genotypes for Field Resistance Against Stem And Root Rot Caused by Macrophomina Phaseolina (Tassi) Goid.
- Correlation Analysis Among Yield and Its Component Traits in F2 Population of Groundnut (Arachis Hypogaea L.) Derived From TCGs 1694 × Icgv 201179
- Effect of Groundnut Based Millet Intercropping System on Yield And Economics of Groundnut (Arachis Hypogaea L.)

