Segregation Distortion Implications and Analysis in Crop Breeding
0 Views
A. RUPA*, B. RUPESH KUMAR REDDY, R. NARASIMHULU AND M. PRADEEP
Department of Genetics and Plant Breeding, ANGRAU-S.V. Agricultural College,Tirupati-517 502.
ABSTRACT
Mendel’s law of segregation assumes equal transmission of alleles from parent to offspring. However, real-world genetic studies frequently reveal deviations from this expected pattern, a phenomenon known as segregation distortion (SD). This refers to the unequal inheritance of alleles during the formation of gametes or the development of zygotes. Such distortion is commonly observed in segregating populations, including F₂ generations, backcrosses, and recombinant inbred lines (RILs). Initially, researchers attributed these deviations are due to technical errors in genotyping or experimental design. However, it is now widely accepted that segregation distortion is often a result of underlying biological mechanisms. These include gametic selection, zygotic selection, meiotic drive, chromosomal rearrangements, and genetic incompatibility. In plant breeding, segregation distortion pose both challenges and opportunities. It may disrupt the accuracy of marker-assisted selection (MAS), QTL mapping, and genetic linkage analysis, potentially leading to the misidentification of desirable traits. However, if properly identified and analyzed, distorted markers can help uncover important genomic regions involved in reproductive barriers or trait inheritance. Advanced statistical methods such as chi-square analysis, likelihood ratio tests and bayesian models as well as specialized mapping tools can assist in detecting and interpreting segregation distortion. Understanding these patterns enables plant breeders to refine selection strategies and accelerate the development of improved crop cultivars.
KEYWORDS: Gametic selection, genomic regions, meiotic drive, marker-assisted selection (MAS), segregation distortion, zygotic selection.
INTRODUCTION
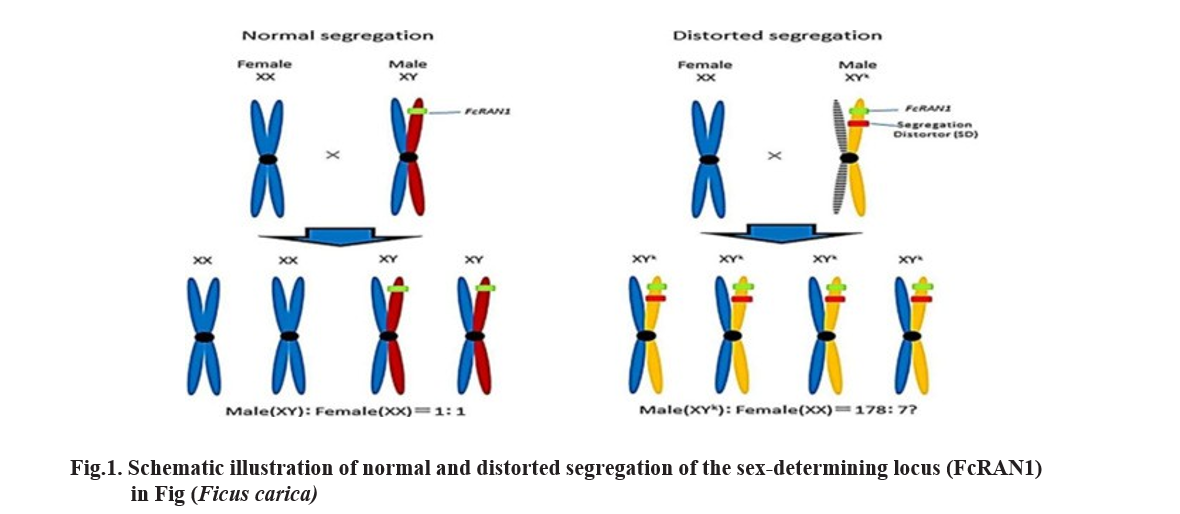
Deviation of the observed genotypic frequencies from the expected mendelian ratios indicates a violation of the law of segregation. Such deviation is referred to as segregation distortion (SD) which was first reported by Mangelsdorf and Jones in 1926 in Maize. SD act as an evolutionary force by influencing the frequency of alleles by manipulating the transmission of alleles during the meiosis. It can increase the frequency of heterozygous alleles through the process called Transmission Ratio Distortion (Huang et al., 2013). It occurs when the genetic distortion affects the transmission of alleles from one generation to the next generation, leading to unequal ratio of different alleles in the offspring. So it is also called as Biased Transmission Distortion. Under normal segregation, the ratio of male (XY) to female (XX) offspring is expected to be 1:1.
Under distorted segregation, due to the presence of a segregation distorter (SD) linked to the FcRAN1 locus, the transmission of the Y chromosome is favored, resulting in a skewed ratio of male (XY⁺): female (XX) = 178:7. Colored bars represent homologous chromosomes carrying the FcRAN1 locus, with red and blue indicating normal alleles and yellow showing the distorted allele carrying SD (Figure:1) (Ikegami et al., 2021). The FcRAN1 genotype shows a strong association with the sex phenotype, such genotypic distortion may affect the evolution of sexual reproduction and sex chromosomes in multiple ways (Kozielska et al., 2010). This is due to SD acts on Y chromosome increasing its transmission to the offspring. The Y chromosome is transmitted to the offspring more frequently than the X. As a result, the frequency of the XY genotype increases in the next generation. By the selfish genetic elements present on the chromosome can manipulate the segregation of chromosomes during meiosis, leading to an increase in their own frequency in the population. It can complicate genetic mapping, QTL analysis and breeding programmes. In Figure 1, the X and Y chromosomes are homomorphic, showing no significant molecular structural differences (Storey, 1975).
Reasons of segregation distortion Marker-associated factors
SD can result from errors at specific marker loci, leading to incorrect genotyping and biased segregation patterns. Marker genotyping errors, such as
contamination, incomplete DNA digestion, or technical failures, as well as the use of inappropriate statistical tests, can produce inaccurate allele scoring and artificially distorted segregation patterns (Diouf & Mergeai, 2012). Residual heterozygosity in parental lines may also contribute to SD when multiple alleles exist at a locus, leading to deviations in the progeny (Diouf & Mergeai, 2012). Additionally, mutations within the primer binding sites of molecular markers can prevent amplification of specific alleles. For instance, a heterozygous plant (A/B) may appear homozygous (A/A) if the B allele fails to amplify, resulting in segregation distortion (Diouf & Mergeai, 2012).
Gametic selection
Selective processes during gamete formation or fertilization can also produce SD. Certain alleles may cause pollen abortion, reducing the frequency of affected alleles in functional pollen and skewing segregation ratios in the progeny (Xu et al., 2013). Alleles that influence the rate of pollen tube growth can affect fertilization success, as faster-growing pollen tubes are more likely to reach and fertilize the ovule, leading to unequal representation of alleles. Similarly, comparative fertilization, where interactions between pollen and pistil tissues preferentially favor certain genotypes, can further contribute to SD (Kim et al., 2014).
Zygotic Selection
Post-fertilization mechanisms can also induce SD. Certain zygotes may be selectively eliminated or
favored, causing deviations from expected Mendelian ratios (Garavello & Pardo, 2020). Additionally, asymmetric chromosome movement during meiosis, such as preferential segregation of specific alleles, can result in SD in the progeny.
Mechanisms of segregation distortion Killer protector system
In this system, gametes carrying a killer gene survive only if they also harbor a linked protector gene; gametes lacking the protector are selectively eliminated. A classical example is the S5 locus in rice, which comprises three tightly linked genes—ORF3, ORF4, and ORF5—that interact to determine gamete viability. In Indica rice, ORF3+ (protector) counteracts the toxic effect of ORF5+ (killer), whereas in Japonica, the absence of ORF3+ leads to programmed cell death of gametes containing only the killer gene, resulting in SD. The evolutionary origin of such systems often involves multiple sequential steps, where the balance between killer and protector genes may be established or disrupted, as observed in ancestral populations of the S5 complex under the parallel-sequential divergence model (Ouyang and Zhang, 2013, Ouyang, 2019; Du et al., 2011; Mi et al., 2020 ). Over evolutionary time, suppressor genes may arise that mitigate the killer’s effect, temporarily restoring Mendelian segregation, before new cycles of distortion emerge. These dynamic interactions illustrate how linked genetic elements can bias allele transmission, influence hybrid sterility, and shape population genetics.
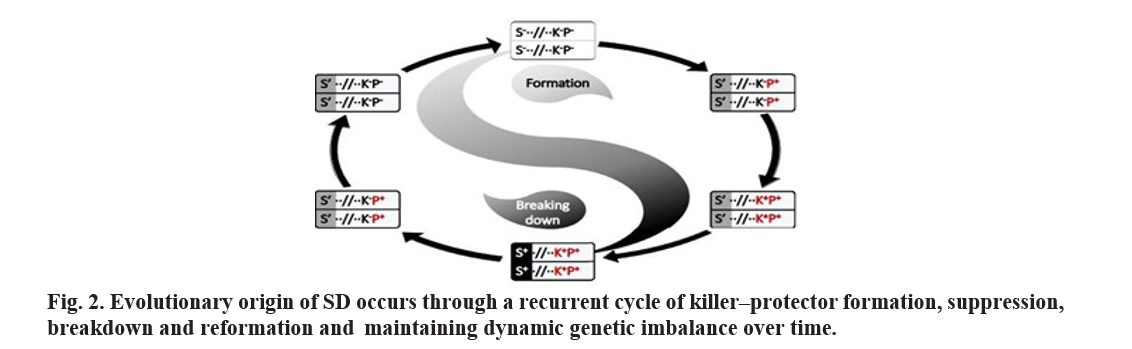
The molecular and evolutionary dynamics of killer–protector systems have been investigated in rice and other species, highlighting their importance in reproductive isolation and breeding strategies (Yang et al., 2012; Xia et al., 2020; Mi et al., 2020).
Meiotic drive
A meiotic driver is a selfish genetic element that manipulates the segregation process to ensure its preferential inclusion in functional gametes of heterozygotes, thereby reducing the transmission of the competing allele. Two ways to be over-represented: gamete killing in males (disruption of alternative sperm) and gonotaxis in females (preferential transmission into the ovule). During spermatogenesis, the driver prevents the formation of functional sperm that do not have a copy of itself. This efficient strategy is referred to as ‘interference’.
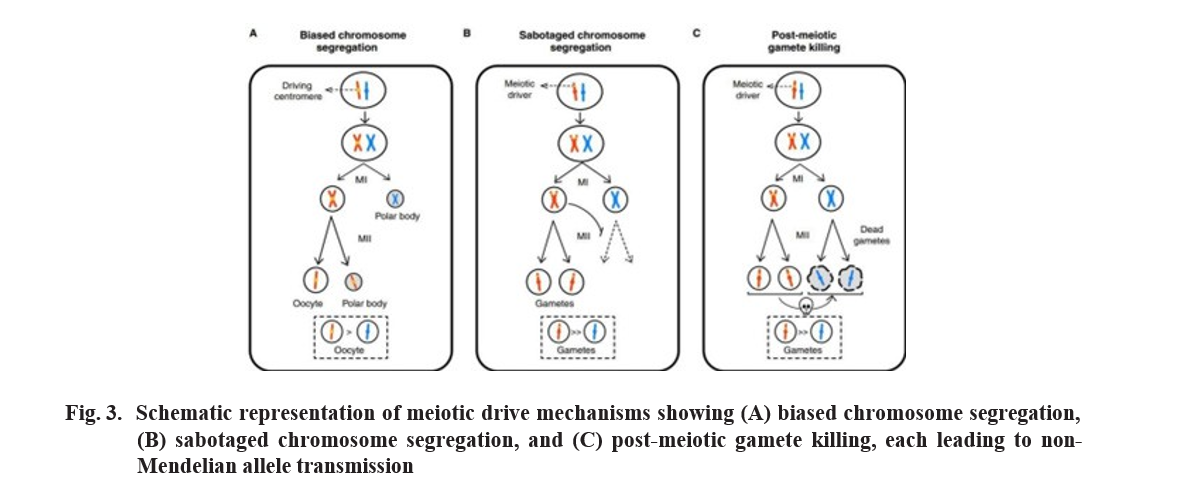
Biased chromosome segregation
Asymmetry of female meiosis where not all meiotic products become gamete. In females, during meiosis where it produces only one large egg cell and small polar bodies that don’t become eggs. Here, one with red have a driver act as a competitor.
Meiotic drivers can act more aggressively during meiosis by sabotaging the ability of competing alleles to complete meiosis. This mechanism is a bit more aggressive. The driver gene (on red chromosome) doesn’t just ensure its own smooth ride. It actively interferes with it’s competitor. Meiosis 1:- occur normally Meiosis 2: Here’s where the, Sabotage happens! the driver gene on the red chromosome disrupts the proper segregation of the competing blue chromosome. It might cause it to break, not sort Correctly.
Post meiotic gamete killing
This is most direct & more aggressive form of meiotic drive. The chromosomes actually segregate(normal). The driver gene then activates a “kill switch”. Driver disrupts gametogenesis and destroy developing gametes carrying the competing allele. This means that after the gametes are formed, any gamete that happen to contain the blue chromosome is actively destroyed.
Fate and evolutionary consequences of gene duplication in the context of segregation distortion
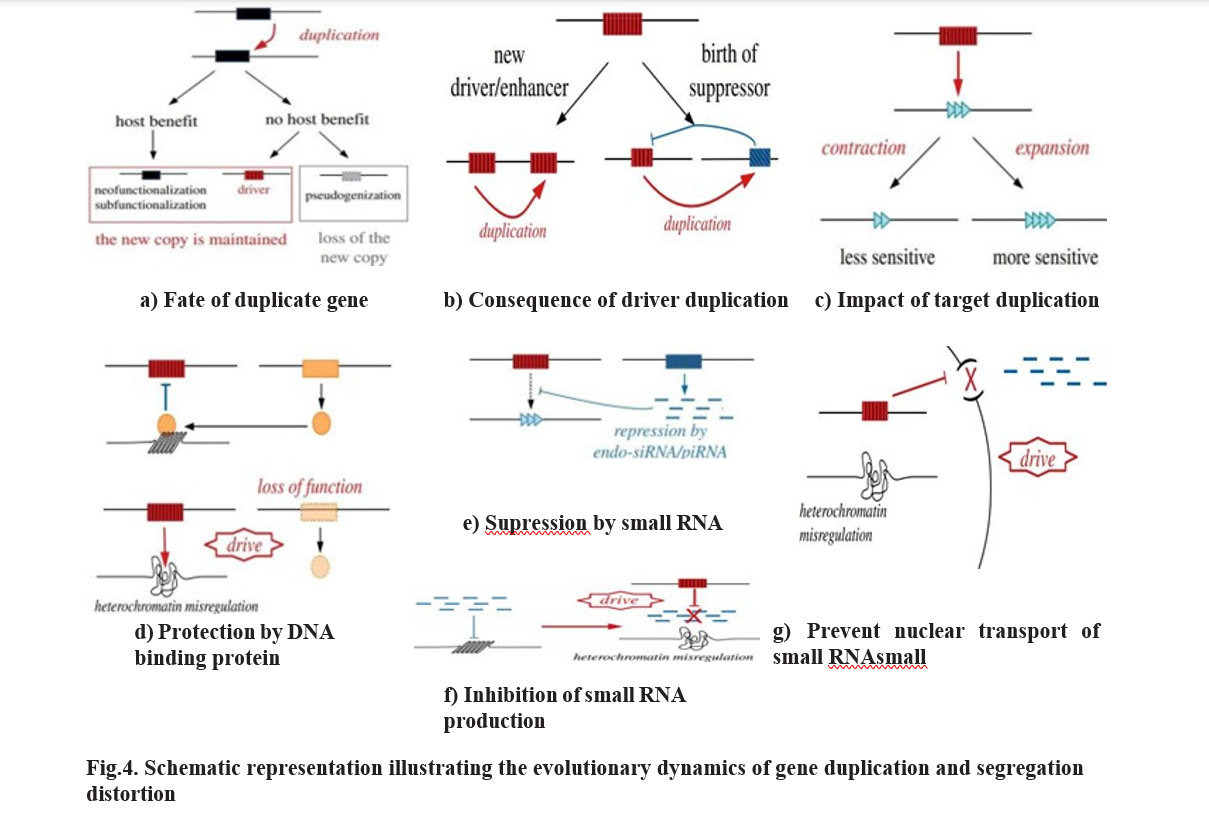
Gene duplication represents a fundamental process driving genome evolution and functional diversification. The evolutionary fate of a newly duplicated gene generally follows two major trajectories depending on whether it benefits the host organism. When the duplicated gene confers a host benefit, it may be retained through neofunctionalization, where the duplicate acquires a novel function distinct from the ancestral copy, or through subfunctionalization, in which ancestral functions are partitioned between the two paralogs (Ohno, 1970; Lynch and Conery, 2000). In some cases, the duplicated copy evolves into a driver gene (figure:4 (a)), a selfish genetic element that distorts Mendelian segregation to favor its own transmission, as reported in Drosophila and Neurospora (Larracuente and Presgraves, 2012). Alternatively, if the new copy provides no host advantage, it typically experiences relaxed selection pressure, leading to degenerative mutations and pseudogenization (Zhang, 2003). However, occasionally such a non-beneficial duplicate may evolve selfish properties, initiating conflict between the host genome and the driver element.
Consequences of driver duplication
When a duplicated driver element further amplifies itself, its segregation distortion activity can intensify, but excessive drive is often deleterious to host fitness. To counterbalance this, the host evolves suppressor genes that mitigate the driver’s effects, restoring fair meiosis. Interestingly, these suppressor genes may themselves arise through secondary duplication events, representing an evolutionary arms race between the driver and its suppressors (Courret et al., 2019; Wedell et al., 2019) (figure: 4(b)).
Impact of target duplication
The interaction between a driver and its target gene can further influence this dynamic. If the target locus becomes less sensitive to the driver’s influence, a contraction in driver prevalence occurs. Conversely, target expansion—through duplication or increased susceptibility—can enhance the driver’s spread (Helleu et al., 2016). Thus, changes in target gene architecture can shape the persistence or decline of segregation distorters (figure: 4(c)).
Host defense mechanisms
Hosts have evolved multiple defense strategies to protect genomic integrity from driver activity. One mechanism involves DNA-binding proteins (figure: 4(d)) that recognize and bind to driver-associated sequences, preventing access to essential chromosomal regions and blocking the driver’s manipulative effects. Another critical mechanism is small RNA-mediated suppression. Host organisms produce endo-siRNAs or piRNAs (figure: 4(e)) that recognize driver transcripts and silence them either transcriptionally or post-transcriptionally (Aravin et al., 2007). However, drivers may counteract this repression by inducing heterochromatin misregulation, (figure: 4(f)) disrupting the host’s small RNA biogenesis and creating a self-reinforcing feedback loop that complicates suppression (Chakraborty et al., 2022). In some cases, the driver induces chromatin barriers that impede nuclear transport of small RNAs, preventing them from entering the nucleus where the driver operates (figure:4 (g)). This spatial barrier enables the driver to escape silencing, perpetuating the genomic conflict.
Ab10 chromosomal meiotic drive in maize: The Rhoades Model
The Abnormal chromosome 10 (Ab10) system in maize (Zea mays) represents one of the most well-characterized examples of chromosomal meiotic drive, first described by Rhoades (1952). The Ab10 haplotype carries large heterochromatic regions, known as knobs, composed primarily of Knob180 and TR-1 tandem repeats. A key component of the drive mechanism is a novel kinesin motor protein, encoded by the Kindr gene, which is essential for activating these knob sequences (Dawe et al., 2018). Upon activation, Kindr binds specifically to the Knob180 and TR-1 repeats, converting them into neocentromeres—additional microtubule attachment sites capable of generating poleward movement during meiosis (Yu et al., 1997; Swentowsky et al., 2020). During anaphase I of meiosis, the Ab10 chromosome exhibits enhanced traction as it is pulled towards the pole by both its primary centromere and the neocentromeric knobs. In contrast, the normal chromosome 10 (N10) lacks these activated knob regions and thus experiences weaker spindle attachment (Hiatt and Dawe, 2003). This differential movement leads to biased segregation, where the Ab10 chromosome is preferentially transmitted to more than 50 per cent of gametes, violating Mendelian expectations. In asymmetric female meiosis, this bias is particularly evident, as the Ab10-bearing chromatid is more frequently incorporated into the functional egg cell rather than the polar bodies, resulting in unequal gamete distribution. Collectively, the Rhoades model demonstrates how structural and molecular innovations—specifically neocentromere formation driven by the Kindr kinesin—allow Ab10 to manipulate the meiotic process for its own transmission advantage (Rhoades, 1952; Dawe et al., 2018; Swentowsky et al., 2020).
Chromosomal abnormalities and their role in segregation distortion
Chromosomal abnormalities represent key structural or numerical alterations that can significantly affect meiotic behavior and genetic transmission. Deletions involve the loss of a chromosomal segment, often including the centromere, which results in the formation of acentric fragments that fail to attach to spindle fibers and are subsequently lost during meiosis (Griffiths et al., 2000; Brown, 2002). In contrast, duplications lead to the doubling of a chromosomal segment, increasing gene dosage and sometimes generating driver loci capable of distorting normal segregation patterns during gametogenesis (Larracuente and Presgraves, 2012). Inversions, which occur when a chromosomal segment is excised and reinserted in the reverse orientation, can interfere with recombination; the resulting acentric and dicentric fragments often fail to segregate properly, leading to gametic loss or sterility (Kirkpatrick, 2010).
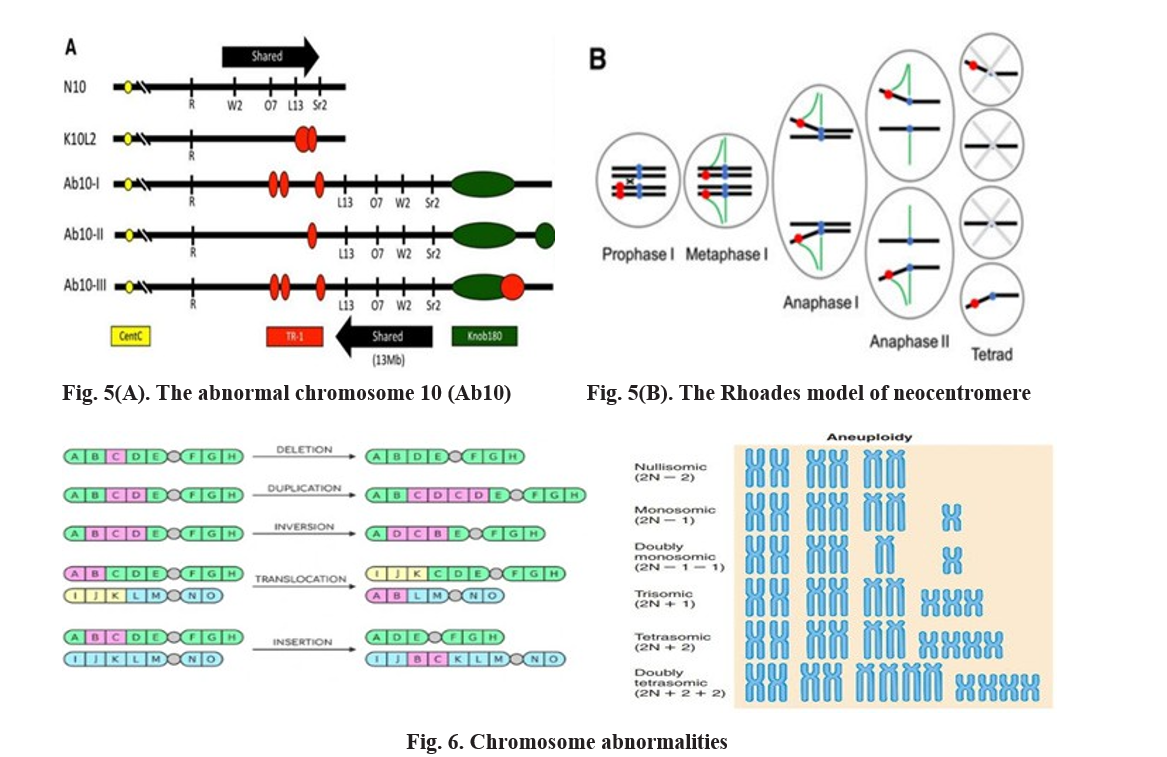
Translocations, involving the relocation of chromosomal segments or entire gene blocks to new positions within the genome, disrupt normal pairing and segregation during meiosis and can consequently influence fertility and segregation ratios (Kumari and Kumar, 2015). Numerical chromosomal changes, including aneuploidy, arise from errors in chromosome segregation, such as nondisjunction, leading to the gain or loss of specific chromosomes.
Such irregularities can result in gametes with unbalanced chromosome numbers—trisomic or monosomic—which, when involved in fertilization, produce offspring with abnormal karyotypes. These numerical anomalies disturb normal Mendelian segregation and contribute to segregation distortion or meiotic drive phenomena, where certain alleles or chromosomes are preferentially transmitted to the next generation (Pardo-Manuel de Villena and Sapienza, 2001; Fishman and Saunders, 2008). Thus, both structural and numerical chromosomal abnormalities play a significant role in shaping genome evolution and reproductive outcomes through their effects on meiotic segregation.
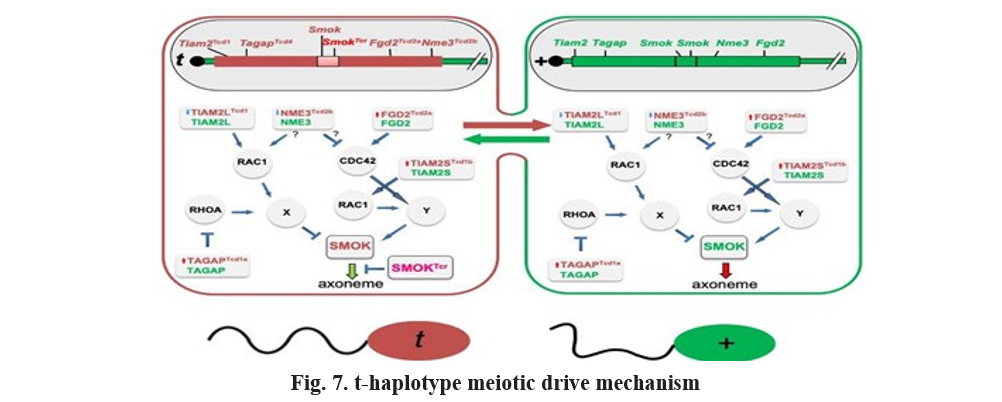
Gametic selection: The t-Haplotype and meiotic drive in mice
The t-haplotype in Mus musculus represents one of the most well-studied examples of a naturally occurring meiotic drive system. It is a large chromosomal region, approximately 40 megabases in length, located on chromosome 17, and encompasses a set of tightly linked genes that collectively promote biased transmission of the t allele (Amaral and Herrmann, 2021). Male mice heterozygous for the t-haplotype (T/t) normally would transmit either the wild-type (T) or t allele to their offspring in equal proportions. However, due to the action of this meiotic drive system, the t allele is transmitted to more than 50% of progeny, demonstrating a clear deviation from Mendelian segregation (Herrmann et al., 1999; Lyon, 2003). The t-haplotype contains multiple genes classified as distorters and a single responder, which function together to manipulate sperm motility.
The distorter genes—such as Tagap, Fgd2, Nme3, and Tiam2—disrupt the normal regulation of the SMOK (Sperm Motility Kinase) protein, a critical component in controlling the directional motility of sperm flagella (Bauer et al., 2012; Herrmann and Bauer, 2012). This disruption impairs the motility of sperm carrying the wild-type T allele, effectively crippling their ability to swim towards the egg. In contrast, sperm carrying the t-haplotype remain functional because of the t-responder (SmokTcr), which compensates for the distorter-induced defects and restores proper motility (Herrmann et al., 1999; Amaral and Herrmann, 2021). This intricate interaction between distorters and the responder within the t-haplotype creates a “selfish” genetic element that biases sperm function and inheritance in its favor, ensuring that t-bearing sperm are more likely to fertilize the egg. Consequently, this mechanism exemplifies how genetic conflicts within the genome can drive evolutionary processes by promoting non-Mendelian inheritance patterns.
Pollen competition and zygotic selection
Pollen competition and zygotic selection represent key post-meiotic mechanisms contributing to segregation distortion in plants. Unilateral incompatibility often requirements of pollen tube growth frequently exceed the pollen’s intrinsic reserves. Furthermore, successful fertilization depends on species-specific signaling between the pollen and the ovule, highlighting the intricate molecular communication underlying reproductive compatibility and selection. Zygotic selection, on the other hand, occurs through the nonrandom loss of diploid zygotes after fertilization. This process can take place during various stages of development, including embryo or seed maturation within the maternal tissue, early germination, or during later juvenile stages (Fishman et al., 2019). Maternal plants that mature only a subset of fertilized eggs can selectively replace less viable zygotes with those carrying more adaptive genotypes, promoting offspring fitness. Two major factors contributing to zygotic selection include environmental selection, where specific germination or growth conditions favor one parental genotype, and inbreeding depression, which promotes segregation distortion at single loci through the expression of lethal or deleterious recessive alleles in homozygous individuals. Collectively, these mechanisms demonstrate how post-fertilization processes can shape genetic transmission patterns and influence evolutionary outcomes in plant populations.
leads to the stylar rejection of pollen grains carrying incompatible or mismatched genotypes, thereby generating pollen-specific, style-dependent segregation distortion (SD) (Fishman et al., 2019). Pollen tube performance and successful fertilization are highly influenced by interactions between the male gametophyte and female reproductive tissues, as the energetic
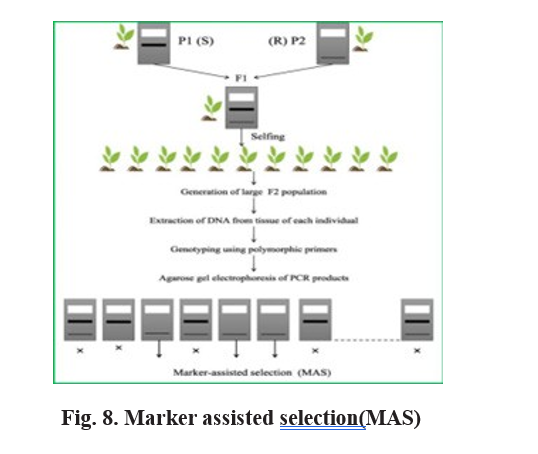
Implications of segregation distortion in crop breeding
Segregation distortion (SD) poses significant challenges and opportunities in modern crop breeding programs by influencing the predictable transmission of alleles. Since SD results in the non-Mendelian inheritance of certain alleles, it complicates the prediction of offspring genotypes and the selection of desirable traits, as alleles may be passed on at frequencies deviating from the expected 1:1 ratio (Liu et al., 2010; Xu, 2008). This distortion directly affects marker-assisted selection (MAS), where molecular markers are used to track genes or quantitative trait loci (QTLs). Skewed allele transmission can weaken or misrepresent marker–trait associations, leading to unreliable identification of genomic regions controlling agronomically important traits (Tao et al., 2002). In genetic mapping, the presence of SD interferes with the accuracy of linkage analysis by distorting recombination frequency estimates, there by reducing the precision of QTL mapping and fine-mapping efforts (Xu, 2008). Moreover, if the distorted segregation pattern disproportionately favors undesirable alleles, breeding efficiency declines, increasing the time and resources required to achieve breeding objectives such as enhanced yield, stress tolerance, or disease resistance. Despite these challenges, segregation distortion also offers valuable biological insights into the mechanisms of gametic selection, meiotic drive, and reproductive incompatibility—processes that, once understood, can be exploited to manipulate genetic inheritance and improve breeding outcomes (Fishman and Saunders, 2008; McDermott and Noor, 2010).
Furthermore, SD influences gene flow and population structure by altering allele frequencies among breeding lines or populations, potentially reducing genetic diversity or causing unintended introgression effects in hybrid programs (Lu et al., 2002). Therefore, recognizing and managing segregation distortion is essential for breeders to refine selection strategies, maintain genetic balance, and better utilize the full genetic potential of crop germplasm.
Approaches for the Analysis of segregation distortion
Segregation distortion (SD) refers to the deviation of observed genotypic ratios from the expected Mendelian inheritance patterns. To understand and quantify this deviation, several statistical and computational methods have been developed, ranging from classical hypothesis-testing procedures to advanced probabilistic and mixed-model approaches (Xu, 2008). The Chi-square test is the most widely used and straightforward statistical method for detecting segregation distortion. It evaluates whether the observed frequencies of alleles or genotypes conform to the expected Mendelian ratios. Under thenull hypothesis (H₀), segregation follows Mendelian expectations, while under the alternative hypothesis (H₁), significant deviation occurs. The Chi-square statistic is computed and compared to the critical value (3.84 at p = 0.05, df = 1). If the calculated value exceeds this threshold, the null hypothesis is rejected, indicating significant segregation distortion in the analyzed population. The Likelihood Ratio Test (LRT) provides a more powerful and flexible alternative to the Chi-square approach, particularly when dealing with small sample sizes or complex inheritance patterns. It compares the likelihood of the observed data under both null and alternative hypotheses to determine the presence of distortion. The test statistic follows a Chi-square distribution, and a significant result suggests deviation from Mendelian segregation ( Li et al., 2007).
The LRT is especially useful in mapping populations where segregation distortion may vary among loci or chromosomes.
The Bayesian segregation ratio analysis extends traditional inference by incorporating prior information with observed data to estimate posterior probabilities of segregation ratios. This approach uses Bayes’ theorem to update prior expectations—typically the Mendelian 3:1 or 1:2:1 ratios—based on observed genotype frequencies. If the posterior probability strongly supports a non-Mendelian ratio, it indicates segregation distortion. Bayesian approaches are advantageous in accounting for uncertainty, small sample sizes, and incomplete datasets (Wang et al., 2019).
Beyond these classical tests, advanced statistical frameworks such as the Expectation–Maximization (EM) algorithm and Generalized Linear Mixed Models (GLMMs) are applied to improve parameter estimation under complex segregation patterns. The EM algorithm iteratively estimates missing or hidden genotypic data, providing more accurate detection of distortion, while GLMMs allow the incorporation of random effects and complex experimental designs (Xu 2008). Quantitative Trait Locus (QTL) mapping is another key analytical approach, used to locate chromosomal regions associated with segregation distortion or traits influenced by distorted loci. QTL mapping involves the selection of an appropriate mapping population, measurement of target traits, genotyping using molecular markers, and statistical association analysis between marker loci and trait variation. This approach helps identify genomic regions showing distortion and their potential impact on quantitative traits (Zhao et al., 2013; Li et al., 2007). A variety of bioinformatics tools and software packages have been developed to facilitate segregation distortion and QTL analysis. Widely used programs include JoinMap, MapQTL, MSTMap, R/qtl, TASSEL, PLINK, PolymapR, GATK, and the CIMMYT Fieldbook. These platforms support data visualization, statistical testing, and marker-trait association studies, offering integrated workflows for segregation and linkage analysis (Van Ooijen, 2011). Together, these statistical, Bayesian, and computational frameworks provide a comprehensive foundation for studying segregation distortion and its genetic consequences in plant breeding and genomics.
A phenomenon known as segregation distortion (SD) refers to the unequal inheritance of alleles during the formation of gametes or the development of zygotes which is commonly observed in segregating populations, including F₂ generations, backcrosses, and recombinant inbred lines (RILs). At the molecular level, the S1TPR locus demonstrates dual roles as both a segregation distorter and protector, a mechanism distinct from conventional killer–protector systems. The S1TPR gene rescues S1-g gametes while selectively eliminating S1-s gametes lacking the protective allele, thereby ensuring biased transmission in favor of the functional allele. In practical breeding applications, incorporating distorted markers has proven beneficial for improving linkage map accuracy and genome coverage. These markers facilitate better grouping of loci within chromosomes and enhance consistency between linkage and physical maps (Xu, 2008). Understanding and accounting for SD in quantitative trait locus (QTL) mapping and marker-assisted selection (MAS) can thus prevent the loss of desirable alleles and improve breeding precision. Ultimately, the success of modern crop breeding programs depends on a thorough understanding of segregation distortion mechanisms. By identifying and managing segregation distortion loci (SDLs), breeders can optimize parent selection, minimize transmission bias, and achieve greater control over the inheritance of favorable traits. This knowledge provides an invaluable foundation for advancing crop genetic improvement and ensuring sustainable breeding outcomes.
LITERATURE CITED
Amaral, A.J. and Herrmann, B.G. (2021) A molecular basis for transmission ratio distortion and meiotic drive in the mouse t-haplotype, Annual Review of Genetics. 55: 165–190.
Aravin, A.A., Hannon, G.J. and Brennecke, J. (2007) The PiRNA pathway in animal germline defense, Science. 318(5848): 761–764.
Bauer, H., Willert, J., Koschorz, B. and Herrmann, B.G. (2012) The t-complex-encoded guanine nucleotide exchange factor Fgd2 is involved in control of sperm motility, Development. 139(17): 3135–3145. Brown, T.A. (2002) Genomes. 2nd ed. New York: Garland Science.
Chakraborty, M. et al. (2022) Evolutionary conflict and chromatin misregulation by meiotic drivers, Nature Communications. 13: 459.
Courret, C., Chang, C.H., Wei, K.H.C., Montchamp-Moreau, C. and Larracuente, A.M., 2019. Meiotic drive mechanisms: lessons from Drosophila, Proceedings of the Royal Society B. 286(1913):
20191430
Dawe, R.K., Lowry, E.G., Gent, J.I., Stitzer, M.C., Swentowsky, K.W., Higgins, D.M., Ross-Ibarra,
- and Kanizay, L.B. (2018) A kinesin drives the segregation distortion of maize Ab10 meiotic drive, Science, 362(6413): 1300–1302.
Diouf, F. B. H., & Mergeai, G., 2012. Segregation distortions and their consequences for plant breeding: A review, Biotechnology, Agronomy, Society and Environment. 16(3): 163–169.
Du HY, Ouyang YD, Zhang CJ, Zhang QF (2011) Complex evolution of S5, a major reproductive barrier regulator, in the cultivated rice Oryza sativa and its wild relatives, New Phytol. 191: 275–287.
Fishman, L. and McIntosh, M., 2019. Standard deviations: the biological bases of transmission ratio distortion, Annual Review of Genetics. 53(1): 347-372.
Fishman, L. and Saunders, A. (2008) Centromere-associated female meiotic drive entails male fitness costs in monkeyflowers, Science. 322(5907): 1559– 1562.
Garavello, M., & Pardo, J., 2020. Analysis of crossover events and allele segregation in the tomato genome. Frontiers in Plant Science. 11: 615. https://doi. org/10.3389/fpls.2020.00615.
Griffiths, A.J., 2005. An introduction to genetic analysis. Macmillan.
Helleu, Q., Gérard, P.R. and Montchamp-Moreau, C. (2016) Sex chromosome drive, Cold Spring Harbor Perspectives in Biology. 8(3). a023143.
Herrmann, B.G. and Bauer, H. (2012) The mouse t-haplotype: A selfish genetic element causing transmission ratio distortion and male sterility, Cold Spring Harbor Symposia on Quantitative Biology. 77: 25–35.
Herrmann, B.G., Koschorz, B., Wertz, K., McLaughlin, K.J. and Kispert, A. (1999) A protein kinase encoded by the t-complex responder gene causes non-Mendelian inheritance, Nature. 402(6758): 141–146.
Hiatt, E.N. and Dawe, R.K. (2003) Four loci on abnormal 10 in maize contribute to meiotic drive of knobs, Genetics. 164(2): 699–709.
Huang, L.O., Labbe, A. and Infante-Rivard, C., 2013. Transmission ratio distortion: review of concept and implications for genetic association studies, Human Genetics. 132(3): 245-263.
Ikegami, H., Shirasawa, K., Yakushiji, H., Yabe, S., Sato, M., Hayashi, T., Tashiro, K. and Nogata, H., 2021. Analysis of the segregation distortion of FcRAN1 genotypes based on whole-genome resequencing of Fig (Ficus carica L.) breeding parents, Frontiers in plant science. 12: 647599.
Kim, B., Lee, J., & Lee, S., 2014. Analysis of segregation distortion and its relationship to hybrid sterility in rice, Rice. 7(1): 3. https://doi.org/10.1186/s12284- 014-0003-8.
Kirkpatrick, M. (2010) How and why chromosome inversions evolve, PLoS Biology, 8(9): e1000501.
Kozielska, M., Weissing, F. J., Beukeboom, L. W., and Pen, I. (2010). Segregation distortion and the evolution of sex-determining mechanisms, Heredity. 104: 100–112. doi: 10.1038/hdy.2009.104
Kumari, S. and Kumar, R. (2015) Chromosomal translocations and their role in genome evolution and speciation, Journal of Cytology & Genetics. 16(1): 1–10.
Mangelsdorf, P.C. and Jones, D., 1926. The expression of Mendelian factors in the gametophyte of maize. Genetics, 11(5): 423.
Larracuente, A.M. and Presgraves, D.C. (2012) The selfish segregation distorter gene complex of Drosophila melanogaster, Genetics. 192(1): 33–53.
Li, W., Lin, Z. and Zhang, X., 2007. A novel segregation distortion in intraspecific population of Asian cotton (Gossypium arboretum L.) detected by molecular markers, Journal of genetics and genomics. 34(7): 634-640.
Liu, G., Yang, J., Xu, H., Hayat, K., Zhang, X. and Guo, H. (2010) Segregation distortion in plants: implications for crop breeding, Euphytica. 174(3): 363–373
Lu, H., Romero-Severson, J. and Bernardo, R. (2002) Chromosomal regions associated with segregation distortion in maize, Theoretical and Applied Genetics. 105(4): 622–628.
Lynch, M. and Conery, J.S. (2000) The evolutionary fate and consequences of duplicate genes, Science. 290(5494): 1151–1155.
Lyon, M.F. (2003) Transmission ratio distortion in mice, Annual Review of Genetics.37: 393–408.
McDermott, S.R. and Noor, M.A.F. (2010) The role of
meiotic drive in hybrid male sterility, Philosophical Transactions of the Royal Society B: Biological Sciences. 365(1544): 1265–1272.
Mi JM, Li GW, Xu CH, Yang JY, Yu HH, Wang GW,
Li XH, Xiao JH, Song HZ, Zhang QF, Ouyang YD (2020) Artificial selection in domestication and breeding prevents speciation in rice, Mol Plant.13: 650–657.
Ohno, S., 2013. Evolution by gene duplication, Springer Science & Business Media.
Ouyang YD (2019) Understanding and breaking down the reproductive barrier between Asian and African cultivated rice: a new start for hybrid rice breeding, Sci China Life Sci. 62: 1114–1116.
Ouyang YD, Zhang QF (2013) Understanding reproductive isolation based on the rice model, Annu Rev Plant Biol 64: 111–135
Pardo-Manuel de Villena, F. and Sapienza, C. (2001) Female meiosis drives karyotypic evolution in mammals, Genetics.159(3): 1179–1189.
Rhoades, M.M. (1952) Preferential segregation in maize.
Genetics. 37(4): 406–424. Storey, W. B. (1975). “Figs,” in Advances in Fruit Breeding, eds J. Janick and J. N. Moore (West Lafayette, IN: Purdue University Press), 568–589.
Swentowsky, K.W., Gent, J.I., Lowry, E.G., Higgins, D.M. and Dawe, R.K. (2020) TR-1 tandem repeats create the neocentromere activity associated with the maize Ab10 meiotic drive system, Genetics. 216(2): 469–480.
Tao, Q., Wang, A., Zhang, L., Xu, J. and Zhang, J. (2002) Segregation distortion in rice and its effect on mapping accuracy, Theoretical and Applied Genetics. 104(3): 476–483.
Van Ooijen, J.W., 2011. JoinMap® 4.1, software for the calculation of genetic linkage maps. Kyazma B.V., Wageningen, Netherlands.
Wang, J., Wang, W. and Li, H., 2019. Bayesian approaches for the detection of segregation distortion loci in plant populations, BMC Genetics. 20(1): 56.
Wedell, N., Price, T.A.R. and Lindholm, A.K. (2019) Meiotic drive and the evolution of female multiple mating, Evolution Letters. 3(3): 265–279.
Xia, F. and Ouyang, Y., 2020. Recurrent breakdown and rebalance of segregation distortion in the genomes: battle for the transmission advantage. Abiotech. 1(4): 246-254.
Xu, S., 2008. Quantitative trait locus mapping can benefit from segregation distortion, Genetics. 180(4) : 2201-2208.
Xu, X., Li, L., Dong, X., Jin, W., & Chen, S., 2013.
Gametophytic and zygotic selection leads to segregation distortion through in vivo induction of a maternal haploid in maize, Journal of Experimental Botany. 64(4): 1083–1096. https://doi.org/10.1093/ jxb/ers393.
Yang JY, Zhao XB, Cheng K, Du HY, Ouyang YD, Chen JJ, Qiu SQ, Huang JY, Jiang YH, Jiang LW, Ding JH, Wang J, Xu CG, Li XH, Zhang QF (2012) A killer-protector system regulates both hybrid sterility and segregation distortion in rice, Science. 337: 1336–1340.
Yu, H.G., Hiatt, E.N., Chan, A., Sweeney, M. and Dawe, R.K. (1997) Neocentromere-mediated chromosome movement in maize, Journal of Cell Biology. 139(4): 831–840.
Zhang, J. (2003) Evolution by gene duplication: an update. Trends in Ecology & Evolution. 18(6): 292–298.
Zhao, K., Aranzana, M.J. and Nordborg, M., 2013. QTL mapping in structured plant populations, Nature Reviews Genetics. 14(11): 879–890.
- Factors Influencing Tribal Women Participation in Van Dhan Vikas Kendra in Alluri Sitarama Raju District in Andhra Pradesh
- Yield and Economics of Greengram Varieties Under Varied Sowing Windows During Rabi Season
- A Study on Profile of Farmers on Drone Technology Application in Agriculture of Rayalaseema Region of Andhra Pradesh
- Studies on Combining Ability for Grain Yield and Its Attributing Traits In Maize (Zea Mays L.)
- Aphidophagous Coccinellid Fauna Associated With Pulses and Oilseed Crop Ecosystems in Tamil Nadu
- Perception and Willingness to Adopt Drone Technology in Tirupati District – A Study on Farmers Profile

