Estimation of Genetic Variability for Yield and Yield Attributing Traits in Sesame (Sesamum Indicum L.)
0 Views
J. ARPITA*, D. BHARATHI, N. SABITHA AND B. RAMANA MURTHY
Department of Genetics and Plant Breeding, S.V. Agricultural College, ANGRAU, Tirupati-517 502.
ABSTRACT
One hundred and four sesame genotypes were evaluated for genetic variability parameters using 18 traits during rabi 2023-24 in dry land farm of S.V. Agricultural College, Tirupati, Andhra Pradesh. Analysis of variance revealed significant differences among the genotypes for all the traits except for 1000 seed weight and capsule length. High estimates of genotypic coefficient of variability (GCV) and phenotypic coefficient of variability (PCV) were recorded for the traits number of secondary branches per plant followed by seed yield per plant and number of capsules per plant. Heritability studies revealed that all the traits except plant height, number of primary branches per plant and capsule length showed high heritability. High heritability coupled with high genetic advance was exhibited by number of secondary branches per plant, seed yield per plant, plant height to first capsule number of capsules per plant, and 1000 seed weight indicating the prevalence of additive gene action in their inheritance.
KEYWORDS: Genetic variability, genetic advance as percent of mean, heritability, sesame.
INTRODUCTION
Sesame (Sesamum indicum L., 2n = 26), the most important prehistoric oilseed crop, belongs to the Pedaliaceae family and serves as a substitute cash crop for small land owners in developing countries due to its excellent adaptability to harsh environments. It is also called as “Queen of oil seeds” due to its high oil content (40 to 63 per cent), mild flavour, and pleasant edible oil. Sesame seeds are abundant in amino acids, proteins, vitamin B, E, minerals, and antioxidants including sesamin, sesamolin and γ-tocopherol. Sesame oil has resistance to oxidative deterioration, and a high level of unsaturated fatty acids (Pusadkar et al., 2015).
India occupies second position in area and production of oilseed crops in the World next to Myanmar. Yield is the most important trait for any crop improvement. Since, yield is a complex trait controlled by many genes, yield can be enhanced by improving the yield contributing traits.
Regardless of the numerous beneficial effects of sesame oil, the productivity is low due to its rainfed cultivation in marginal and sub marginal lands under poor management and input starved conditions, lack of synchronous maturity, shattering, yield instability and lack of high yielding cultivars. Therefore, it is essential to produce high yielding varieties with favourable features in order to overcome the low productivity and yield plateau in sesame.
The success of any crop improvement program depends upon the nature and magnitude of genetic variability present in the crop. The commencement of any systematic breeding programme, requires an understanding of the kind and amount of genetic variability existing in the gene pool. This is because a high level of genetic variability in the base material increases the likelihood of a desired plant type emerging. Keeping this in view, the present investigation was carried out to evaluate the genetic variability parameters among the sesame genotypes for yield and yield attributing traits.
MATERIAL AND METHODS
The present investigation was carried out during rabi 2023-24 on 100 sesame genotypes along with 4 checks collected from Regional Agricultural Research Station, Tirupati, Andhra Pradesh. The experiment was conducted in an unreplicated Augmented Block Design. Each genotype was sown in single row of four meters length with a spacing of 30 cm between rows and 15 cm between the plants. Eighteen yield contributing traits (qualitative and quantitative) were measured in five randomly selected plants from each genotype, to assess the magnitude of variability parameters among the sesame genotypes. The estimation of genetic parameters was done with the help of Grapes software.
Genotypic and phenotypic coefficient of variation
The GCV and PCV were estimated by using the
formula:

GCV and PCV were categorized into Low = Less than 10%; Moderate = 10-20%; High =More than 20%.
Broad sense heritability
The proportion of genotypic variance to the total variance of the population is referred to as heritability in broad sense
![]()
and was calculated by the formula given by Lush (1940).
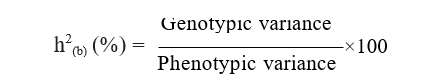
Genetic advance as percentage of mean
Genetic advance as percentage of mean was calculated by the following formula:

GAM was categorized as < 10 per cent as low, 10-20 per cent as moderate, >20 per cent as high.
RESULTS AND DISCUSSION
The analysis of variance revealed significant variation among the genotypes for all the traits except 1000 seed weight and capsule length indicating the presence of considerable genetic variability among the experimental material under study (Table 1). Presence of variability in the initial breeding material ensures better chances of producing a desired genotype.
The study found that the phenotypic coefficient of variation (PCV) surpassed the genotypic coefficient of
variation (GCV) for all traits, highlighting environmental influence on trait expression. This finding aligns with previous findings by Sasipriya et al. (2022). Mean, range, heritability, GCV, PCV and genetic advance as
Table 1. Analysis of variance for yield and yield attributing traits
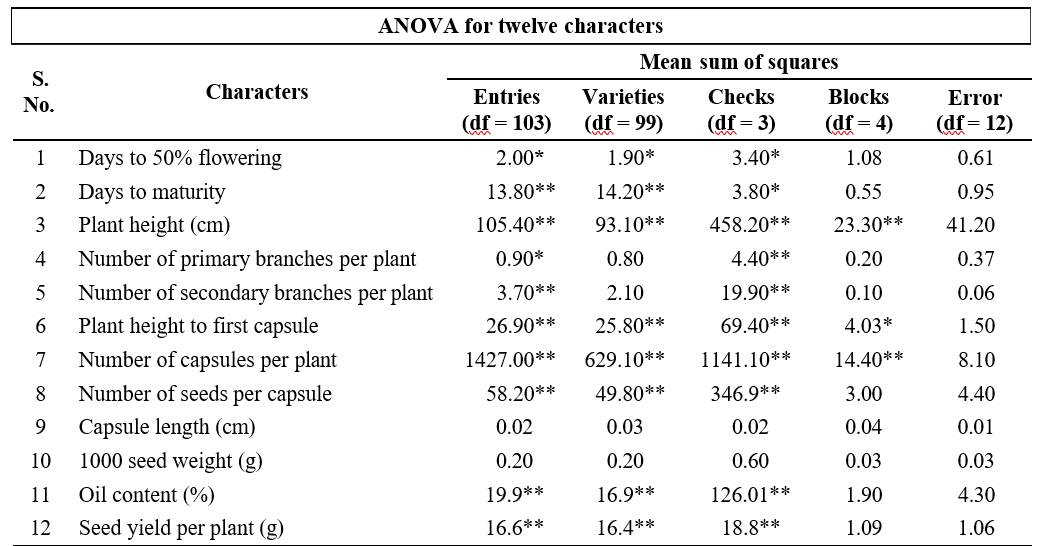
Table 2. Genetic parameters of variability for yield and its attributing traits
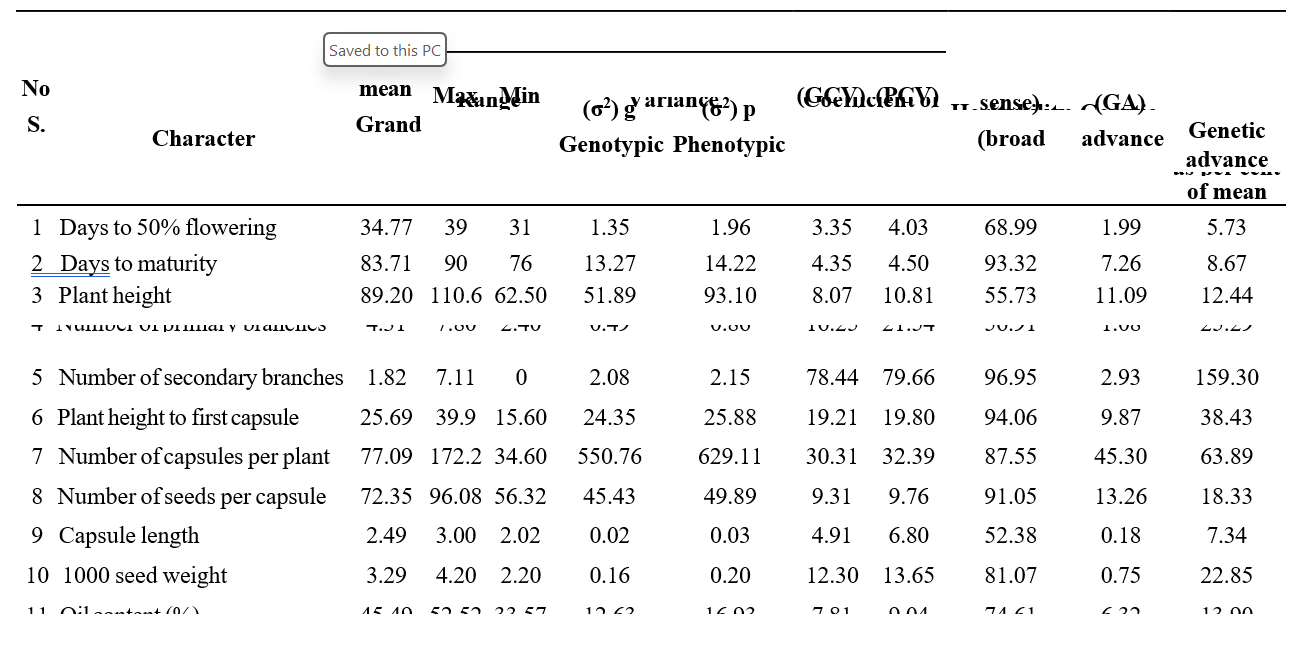
per cent of mean are presented in Table 2. High levels of GCV and PCV were observed for the traits number of secondary branches plant- 1 (GCV: 78.44%; PCV: 79.66%) followed by seed yield plant-1, number of capsules plant-1. This indicates the presence of sufficient amount of variation among the genotypes for these traits. The findings of this study are consistent with the results obtained by Kadvani et al. (2020), Sultana et al. (2019). Moderate levels of GCV and PCV were observed in plant height to first capsule (GCV: 19.21%; PCV: 19.8%) and thousand seed weight (GCV: 12.3%; PCV: 13.65%). The outcomes corroborate those obtained by Kadvani et al. (2020), Kalaiyarasi et al. (2019), Umamaheswari et al. (2019), and Bedawy et al. (2018). The lowest estimates of GCV and PCV were found for the traits number of seeds capsule-1 (GCV: 9.31%; PCV: 9.76%) and oil content (GCV: 7.81%; PCV: 9.04%), followed by capsule length, days to maturity, and days to 50% flowering. The findings align with the conclusions reached by Kadvani et al. (2020), Gautam et al. (2023), Pavani et al. (2020), Kalaiyarasi et al. (2019), Sasipriya et al. (2023).
High estimates of heritability (broad sense) was observed for number of secondary branches plant-1 (96.95%) followed by seed yield plant-1 (95.21%), plant height to first capsule (94.06%), days to maturity (93.32%), number of seeds capsule-1 (91.05%), number of capsules plant-1 (87.55%), 1000 seed weight (81.07%), oil content (74.61%) and days to 50 % flowering (68.99%).
High estimates of genetic advance as percent of mean was recorded for number of secondary branches plant-1 (159.30%) followed by seed yield plant-1 (79.91%), number of capsules per plant (58.5%), plant height to first capsule (38.43%), number of primary branches plant-1 (25.29%) and 1000 seed weight (22.85%). This indicates the predominance of additive gene action and the desired results may be obtained through the simple selection. These results were in agreement with the findings of Pavani et al. (2020) for number of capsules plant-1, plant height to first capsule. High heritability along with high genetic advance as percent of mean was recorded by number of secondary branches plant-1, seed yield plant-1, plant height to first capsule, number of capsules plant-1, and 1000 seed weight. Similar type of results were found by Pavani et al. (2020) for number of capsules plant-1, plant height to first capsule; Kiruthika et al. (2018) for number of secondary branches plant-1, number of capsules plant-1 and 1000 seed weight.
The results of the variability study revealed that high to moderate estimates of GCV, PCV, high heritability and high genetic advance as percent of mean was recorded for the traits number of capsules per plant, seed yield per plant, plant height to first capsule and thousand seed weight This specified the dominance of additive gene action in expression of these traits. Therefore, simple selection might be fruitful for improvement of these traits in segregating generations.
LITERATURE CITED
Bedawy, I.M and Mohamed, N.E. 2018. Phenotypic and genotypic variability in a set of Sesame (Sesamum indicum L.) genotypes. Egyptian Journal of Agronomy. 40(3).
Gautam, S., Mishra, S.P., Gautam, S., Rao, S and Dwivedi, 2023. Morphological and biochemical characters studies in sesame (Sesamum indicum L.) varieties.
Kadvani, G., Patel, J.A., Patel, J.R., Prajapati, K.P and Patel, P.J. 2020. Estimation of genetic variability, heritability and genetic advance for seed yield and its attributes in sesame (Sesamum indicum L.). International Journal of Bio-resource and Stress Management. 11(3).
Kalaiyarasi, R., Lokeshkumar, K., Mohanraj, M., Priyadharshini, A and Rajasekar, R. 2019. Genetic variability parameters for yield and yield related traits in sesame (Sesamum indicum L.). International Journal of Current Microbiology and Applied Sciences. 8(8).
Kiruthika, S., Narayanan, S.L., Parameswari, C., Mini, M.L and Arunachalam, P. 2018. Genetic variability studies for yield and yield components in sesame (Sesamum indicum L.). Electronic journal of plant breeding. 9(4).
Pavani, K., Lal Ahamed, M., Ramana, J.V and Sirisha, A.B.M. 2020. Studies on genetic variability parameters in sesame (Sesamum indicum L.). International Journal of Chemical Studies. 8(4).
Pusadkar, P.P., Kokiladevi, E., Bonde, S.V and Mohite, N.R., 2015. Sesame (Sesamum indicum L.) importance and its high quality seed oil: a review. TRENds in biosciences. 8(15): 3900-3906.
Sasipriya, S., Parimala, K., Balram, M and Eswari,
K.B. 2022. Variability and character association in sesame (Sesamum indicum L.). The Pharma Innovation Journal. 11(1).
Sultana, S., Mahmud, F and Rahim, M.A. 2019. Genetic variability studies for selection of elite germplasm in sesame (Sesamum indicum L.). Agronomski glasnik: Glasilo Hrvatskog agronomskog društva. 81(2): 87-104.
Umamaheswari, S., Suganthi, S., Sathiskumar, P and Kamaraj, A. 2019. Genetic variability, correlation and path analysis in sesame (Sesamum indicum L.).
- Effect of Sowing Window on Nodulation, Yield and Post – Harvest Soil Nutrient Status Under Varied Crop Geometries in Short Duration Pigeonpea (Cajanus Cajan L.)
- Nanotechnology and Its Role in Seed Technology
- Challenges Faced by Agri Startups in Andhra Pradesh
- Constraints of Chcs as Perceived by Farmers in Kurnool District of Andhra Pradesh
- Growth, Yield Attributes and Yield of Fingermillet (Eleusine Coracana L. Gaertn.) as Influenced by Different Levels of Fertilizers and Liquid Biofertilizers
- Consumers’ Buying Behaviour Towards Organic Foods in Retail Outlets of Ananthapuramu City, Andhra Pradesh

